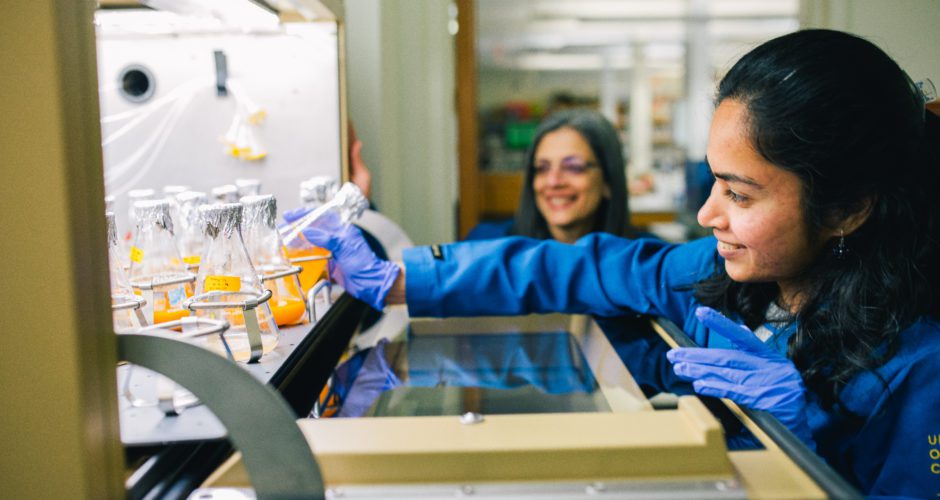
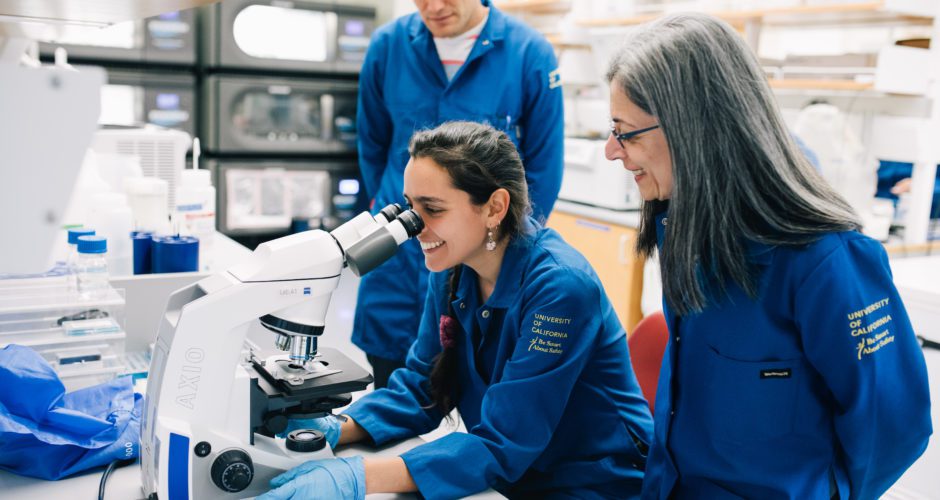
Sabeeha Merchant is a professor of biochemistry, biophysics, and structural biology in the Department of Molecular and Cell Biology and a professor of plant biology in the Department of Plant and Microbial Biology at UC Berkeley, and a faculty senior scientist at Lawrence Berkeley National Laboratory. Her group studies photosynthetic metabolism and metalloenzymes and they use the tools of systems biology and comparative genomics for a big picture perspective. In the early 2000s Merchant joined the team that sequenced the Chlamydomonas genome, work that was recently featured on the Joint Genome Institute’s website. Merchant’s group has now sequenced four other algal genomes. She was elected a member of the US National Academy of Sciences in 2012, the American Academy of Arts and Sciences in 2014 and the Leopoldina in 2016.
QB3-Berkeley: What’s the focus of your lab’s research?
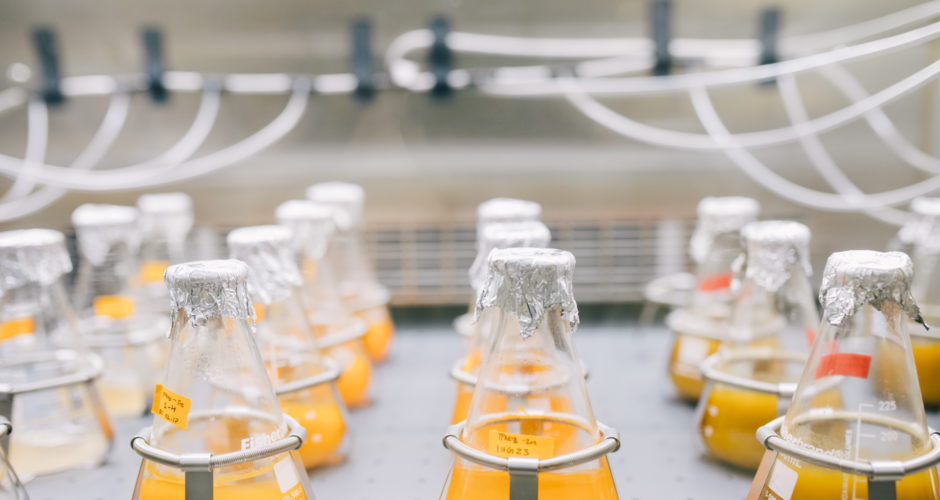
Sabeeha Merchant: We are interested in green algae. Algae are within the plant lineage. We use Chlamydomonas reinhardtii as a reference system to address questions about the metabolism of algae, especially metabolism relating to photosynthesis as well as processes related to how the photosynthesis apparatus is built.

Recently, thanks to genomics, we’ve expanded our experimental base, which makes it possible to essentially use any organism for experimental work or in other words to study any interesting organism. There is an amazing diversity of life on our planet. By looking at how different, but related, organisms have solved a particular problem—e.g. handling a particular nutrient deficient growth environment like Fe deficiency—we can derive general principles in biology.
QB3: What’s the biggest challenge and greatest reward of running your own lab?
SB: There are two challenges for me: one is to identify coworkers who are as passionate about the questions we are asking as I am and the other is to identify sponsors who will support our work.
The reward is also twofold. First, the discoveries we make are rewarding—learning something new that no one else has known previously; in other words, building new knowledge. The other is the products from my laboratory: the students and postdoctoral trainees and scientists who have worked with us and made those discoveries. It’s rewarding to see them learn how to be independent thinkers and learn how to ask and answer questions about nature.
QB3: Are there any forthcoming papers or current projects that you can share with us?
SB: As I mentioned above, our research relies on genomics. At the present time, we have several algal genomics and system biology projects. First, in collaboration with scientists at the Joint Genome Institute we have updated the Chlamydomonas genome and just submitted a paper on the “version 6 assembly.” As I mentioned, Chlamydomonas is a model organism not just for us, but for hundreds of researchers world-wide, especially those who work on photosynthesis, chloroplasts, or the biology of cilia. The genome of Chlamydomonas is therefore a resource for many research groups.
Second, my coworkers Rory Craig, Sean Gallaher, Jeff Moseley and I are developing an alga called Auxenochlorella protothecoides as another reference system for functional studies of green algae. This organism has an interesting metabolic program: it can grow on CO2 and light like other plants and algae, enabling the dissection and study of photosynthesis, and it can also grow on glucose like fungi and animals, in which situation it makes a lot of fat, enabling the dissection and manipulation of metabolism towards producing biodiesel precursors in algae. We have sequenced the genome and we can manipulate genes in this organism, so we are hoping to use this system for producing designer fats. A designer fat created in the lab might serve a medical function or be used in a food; for example, the fats that are used in some infant formulas.
Third, Sean Gallaher and a former coworker Lital Davidi are working on green algae called Dunaliella species to understand how algae handle iron deficiency. Dunaliella is a cosmopolitan alga, and it is also cultivated in farms for production of carotenoids. You can see Dunaliella when you fly through SFO. We have used genomics to deduce metabolic changes in Dunaliella that enable a life on low iron.
QB3: When did you become interested in becoming a scientist?
SB: I am not sure. I started studying science when I was 12 years old. I liked the logic of science. Growing up, we did not have access to science fairs or science projects. It was just classroom knowledge, so it was not until I was an undergraduate in Madison, Wisconsin that I even realized that the knowledge in textbooks was a product of research that someone like me could do.
QB3: What do you like about living in the Bay Area and working at UC Berkeley?
SB: I like the geography, the natural beauty of the area. I like the students on our campus and their commitment to a better future. The staff on campus are professional and the facilities are excellent, both of which support the research enterprise. The proximity to Lawrence Berkeley National Laboratory (LBNL) is a plus for my work because of the direct relevance of our work to the mission of the Department of Energy and hence opportunities to work with LBNL scientists.