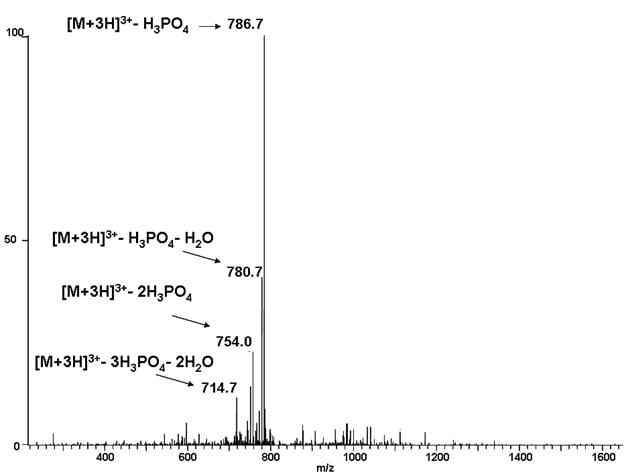
LC MS/MS data can be processed to detect any modifications that may be present in the peptides detected. The modification must add a calculatable molecular weight. Modification analysis can be performed using either one dimensional or multi dimensional “MudPIT” chromatography depending on the sample complexity. We can often find modifications on a target protein even when it is part of a complex mixture. Often modifications of interest can be found simply by searching LC mass spec data collected under standard conditions, but we can do better if the data collection is planned specifically with modification analysis in mind. For some types of modification including phosphorylation, we can collect additional data by CID, HCD or ETD fragmentation that confirms the presence of the modification with higher confidence. Ask us for details.
In some cases, chances of pinpointing the site of a modification increase if overlapping peptides are analyzed or if an enzyme other than trypsin is used to generate peptides. To produce sets of overlapping peptides, we recommend digestion with two or three proteases that vary in their specificity. If there is a suspected site of modification, the proteolytic digest can be planned to produce peptides of optimal mass containing that site. You can use peptidecutter to choose an appropriate protease.
To perform this type of analysis, please refer to the following forms and protocols:
Enzymatic digestion of protein samples
Triple enzymatic digestion of protein samples
Sample desalting for mass spectrometry
We urge you to contact facility staff to discuss your project prior to sample preparation.