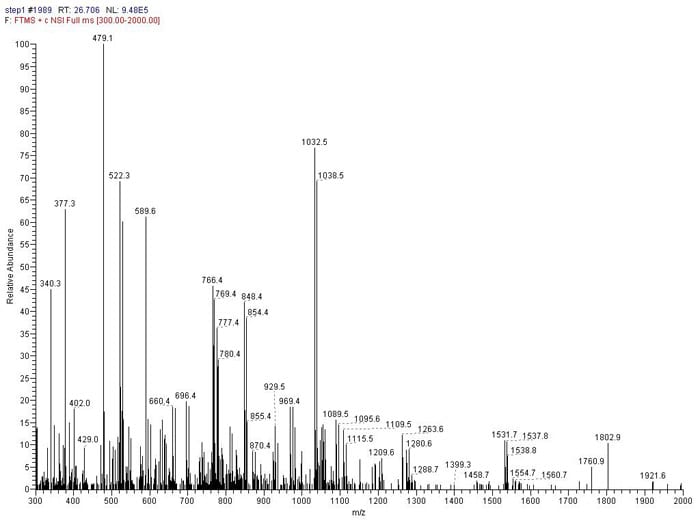
Nanoscale LC MS/MS using CID, HCD or ETD fragmentation allows the identification of individual peptides from a complex mixture. This is a powerful proteomics technique through which the component proteins in a sample can be identified.
Typically, a sample is prepared by tryptic digestion and desalting of a protein mixture obtained in the course of the users’ research. Samples have varied in complexity from purified protein complexes to whole cell extracts.
We load desalted, proteolysed mixture on a nanoscale HPLC column. These columns employ either reverse phase (1-dimensional) or a combination of ion exchange and reverse phase (2-dimensional or mudPIT) separation chemistries. The eluent from the LC column is subjected directly to tandem mass spectrometry, and the mass and fragmentation spectrum of major ions is recorded.
A computer program suite is then used to identify the peptides that gave the spectra that have been collected. It queries a sequence database for the appropriate organism, calculating theoretical fragmentation spectra for all possible peptides and comparing them to the data collected.
The final output for the user is a file listing each gene for which peptides were found in the data. Each peptide is listed along with statistics showing the quality of the data.
One dimensional separations are appropriate for samples where the expected complexity is 1 to 10 proteins. Between 10 ng and 500 ng are needed for the analysis.
Two dimensional “MudPIT” separations are appropriate for more complex mixtures. Between 100 ng and 10 ug are needed for analysis.
Users generally prepare a proteolysed, desalted sample for one dimensional or two dimensional analysis.
To perform this type of analysis, please refer to the following forms and protocols:
Enzymatic digestion of protein samples
Sample desalting for mass spectrometry
We urge you to contact facility staff to discuss your project prior to sample preparation.