Sample Preparation Guidelines
Analysis of intact, non-digested proteins or peptides by mass spectrometry can provide confirmation that a gene product of interest has been biosynthesized and isolated. This measurement can also provide information about the presence of degradation or truncation products, amino acid substitutions, bioconjugation products, and post-translational modifications such as phosphorylation, methylation, glycosylation, etc. Proper sample preparation is crucial for obtaining a high-quality mass spectrum.
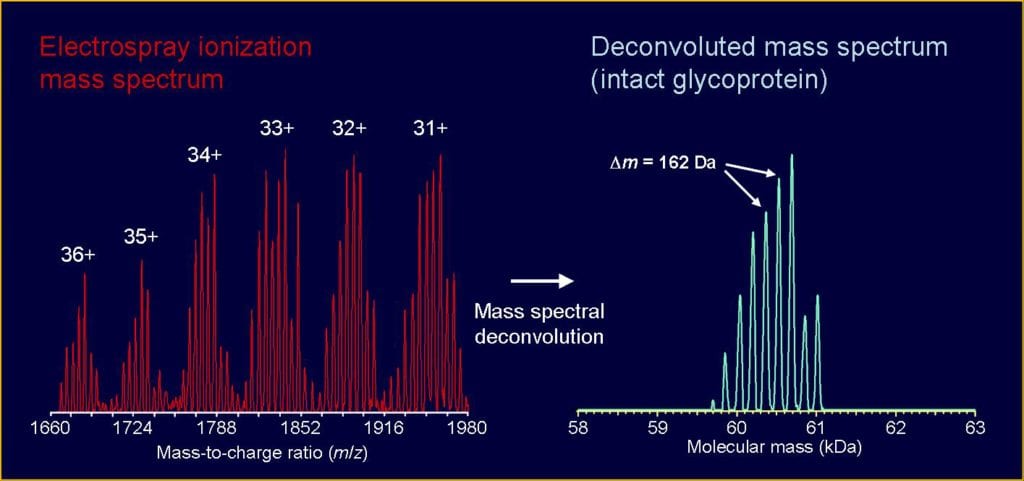 Target analyte concentration: At least 300 picomoles.
Target analyte concentration: At least 300 picomoles.
Typical solution composition: Aqueous (in water).
- Water: Distilled, deionized water.
- Organic solvent: None needed. Glycerol OK (up to 20%).
- Buffers: Common buffers and salts OK, up to 500 millimolar.
Target sample volume: 5 to 100 microlitersInterfering contaminants – Avoid or remove from sample
- Detergents/surfactants: For example, PEG, PPG, Tween, CHAPS, etc.
Target analyte concentration: In the range 3 to 50 pmol/µL
Typical solution composition: Water/organic solvent (1:1) with organic acid (0.1% to 1%)
- Water: Distilled, deionized water.
- Organic solvent: Typically acetonitrile. Other solvents such as methanol, ethanol, n-propanol, isopropanol, etc., will also suffice. The solvent should be HPLC grade. The exact ratio of water to organic solvent is not critical, although 1:1 is typical.
- Organic acid: Formic acid or acetic acid.
Minimum sample volumes
- Syringe infusion electrospray: At least 100 microliters.
- Nanoelectrospray: At least 5 microliters.
Interfering contaminants – Avoid or remove from sample
- Salts
- Metal cations: Li+, Na+, K+, Ca2+, Mg2+, etc
- Inorganic anions such as phosphate, sulfate, and halides
- Alkylammonium salts
- Guanidinium salt
- Detergents/stabilizers: For example, SDS, PEG, PPG, Tween, CHAPS, Triton, and urea.
- Buffers: For example, HEPES, PBS, MES, MOPS, and Tris.
Acceptable buffers – Use only if necessary (at concentrations ≤ 100 mM)
- Ammonium acetate
- Ammonium bicarbonate
- Ammonium formate
Useful sample clean up tools
Guidelines to minimize keratin contamination
- Perform as much of the work as possible within a laminar flow hood or a biological safety cabinet (BSC).
- A clean-room lab coat, nitrile or vinyl gloves and, if necessary, sleeve protectors should be worn.
- The inside of the BSC should be wiped down with water and ethanol prior to beginning work.
- All containers, tools, and apparatus should be wiped down with water and ethanol prior to placing them in the BSC.
- Previously opened containers should not be opened within the BSC.
- Do not use latex gloves or tubing.
Solutions
- 25 mM ammonium bicarbonate (aqueous)
- 25 mM ammonium bicarbonate in 1:1 acetonitrile/water
- 45% water/50% acetonitrile/5% formic acid
- 12.5 ng/µL (525 fmol/µL) trypsin in 25 mM ammonium bicarbonate (aqueous)
Procedure
- Excise the stained bands of interest from the gel using a scalpel or razor blade.
- Dice each gel slice of interest into small pieces (approximately 1 mm 2) using a scalpel or razor blade and add into a 0.65 mL siliconized tube.
- Add enough 25 mM ammonium bicarbonate in 1:1 acetonitrile/water to fully immerse the gel pieces and vortex for 10 minutes.
- Remove the supernatant using a gel-loading pipette tip and discard.
- Repeat steps 3 and 4 up to three times.
- Vacuum centrifuge the gel pieces to complete dryness (~20 to 30 minutes).
- Reduce and alkylate (optional – recommended for disulfide-rich proteins).
- Prepare fresh solutions.
- 10 mM dithiothreitol (DTT) in 25 mM ammonium bicarbonate with 10% acetonitrile
- 55 mM iodoacetamide in 25 mM ammonium bicarbonate (aqueous)
- Add enough 10 mM dithiothreitol solution to fully immerse the gel pieces and vortex and centrifuge briefly.
- Incubate at 56 °C for one hour.
- Cool to room temperature and remove and discard the supernatant.
- Add enough 55 mM iodoacetamide solution to fully immerse the gel pieces and vortex and centrifuge briefly.
- Incubate in the dark at room temperature for 45 minutes.
- Remove and discard the supernatant.
- Add enough aqueous 25 mM ammonium bicarbonate to fully immerse the gel pieces and vortex and centrifuge briefly.
- Remove and discard the supernatant.
- Add enough 25 mM ammonium bicarbonate in 1:1 acetonitrile/water to fully immerse the gel pieces and vortex and centrifuge briefly.
- Remove and discard the supernatant.
- Repeat steps h to k.
- Vacuum centrifuge the gel pieces to complete dryness (~20 to 30 minutes).
- Prepare fresh solutions.
- Add one volume of the trypsin solution. This volume can be estimated from the volume of the excised gel band. For example, 2 mm × 5 mm × 1 mm = 10 mm 3 = 10 µL.
- Incubate on ice or at 4 °C for 30 minutes.
- Remove and discard any excess trypsin solution remaining after 30 minutes.
- If necessary, add a minimum amount of aqueous 25 mM ammonium bicarbonate to keep the gel pieces hydrated during digestion.
- Centrifuge briefly and incubate at 37 °C for 6 to 8 hours.
- Add a volume of water to the digest equal to two or three times the volume of the excised gel piece, vortex for 10 minutes, sonicate for 5 minutes, and centrifuge briefly.
- Extract the supernatant and transfer it into a fresh centrifuge tube (Tube I; do not discard).
- Add enough 45% water/50% acetonitrile/5% formic acid to the tube containing the gel pieces so that the pieces are fully immersed, vortex for 10 minutes, sonicate for 5 minutes, and centrifuge briefly.
- Extract the supernatant and add it into Tube I.
- Repeat steps 15 and 16 twice.
- Reduce the volume of Tube I to 10 µL using a vacuum centrifuge prior to submitting samples for analysis by LC-MS.
References
- Rosenfeld, J.; Capdevielle, J.; Guillemot, J. C.; Ferrara, P. “In-gel digestion of proteins for internal sequence analysis after one- or two-dimensional gel electrophoresis.” Anal. Biochem.1992, 203, 173-179.
- Hellman, U.; Wernstedt, C.; Gonez, J.; Heldin, C.-H. “Improvement of an ‘in-gel’ digestion procedure for the micropreparation of internal protein fragments for amino acid sequencing.” Anal. Biochem.1995, 224, 451-455.
- Shevchenko, A.; Tomas, H.; Havlis, J.; Olsen, J.V.; Mann, M. “In-gel digestion for mass spectrometric characterization of proteins and proteomes.” Nature Protocols.2006, 1, 2856-2860.
- Biringer, R. “Protocol for a keratin-free environment.” Thermo Electron Corp. (accessed September 12, 2007).
Stock solutions
- 100 mM iodoacetamide in water (make fresh, keep from light)
- 1 μg/μl mass spec-grade trypsin in 50 mM acetic acid
- 8 M urea, 50 mM Tris, pH 7.0
- 500 mM DTT, 50 mM Tris, pH 7.0
1. Denature protein. If necessary, reduce cysteines and disulfides with DTT, alkylate all cysteines with iodoacetamide, quench excess iodoacetamide with a bit more DTT.
- Keep pH below 8.5 to reduce alkylation of other nucleophiles.
- To avoid lysine carbamylation, use fresh urea and minimize heating.
2. Digest protein with trypsin.
- Digest >10 μM protein.
- Use 250-500 ng trypsin.
- Keep final free thiol concentration below 10 mM in digestion.
- Keep urea concentration below 1 M in digestion.
- Aim for at least ~50 picomoles digested protein for analysis.
Example: 66 kDa protein with 7 cysteines per monomer:
Denaturation/Reduction:
5 μl 765 μM protein
20 μl 8M urea
0.5 μl 500 mM DTT
25.5 μlIncubate at 55ºC for 20 min
final = 150 μM protein; x7 cysteines = 1.05 mM free thiol; 6.27 M urea; 9.8 mM DTTAlkylation:
+6 μl 100 mM iodoacetamide
31.5 μlIncubate at RT for 30 min in dark
final = 19 mM iodoacetamide, 5.08 M urea, 7.9 mM DTT, 121 μM proteinQuench:
+2 μl 500 mM DTT
33.5 μlIncubate at RT for 20 min; store unused alkylated protein at -20ºC
final = 17.9 mM iodoacetamide, 4.78 M urea, 37.3 mM DTT, 114 μM proteinDigestion:
6 μl alkylated protein
2.5 μl 1M Tris, pH 7.0
0.5 μl 100 mM CaCl2 (if protein is stable with it)
0.5 μl 1 μg/μl trypsin
41.5 μl water
50 μlDigest overnight at RT
final = 13.5 μM protein, 500 ng trypsin, 50 mM Tris pH 7.0, 1 mM Ca2+, 574 mM urea, 4.5 mM DTT, 2.1 mM iodoacetamideAcknowledgment: Dr. Jonathon Winger is gratefully acknowledged for his generous assistance in preparing the protocol.
References
- Rebecchi, K. R.; Go, E. P.; Xu, L.; Woodin, C. L.; Mure, M.; Desaire, H. “A general protease digestion procedure for optimal protein sequence coverage and post-translational modifications analysis of recombinant glycoproteins: application to the characterization of human lysyl oxidase-like 2 glycosylation.” Anal. Chem. 2011, 83, 8484-8491.
- Hervey, W. J., IV; Strader, M. B.; Hurst, G. B. “Comparison of digestion protocols for microgram quantities of enriched protein samples.” J. Proteome Res. 2007, 6, 3054-3061.
- Strader, M. B.; Tabb, D. L.; Hervey, W. J.; Pan, C.; Hurst, G. B. “Efficient and specific trypsin digestion of microgram to nanogram quantities of proteins in organic-aqueous solvent systems.” Anal. Chem. 2006, 78, 125-134.
-
Brownridge, P.; Beynon, R. J. “The importance of the digest: Proteolysis and absolute quantification in proteomics.” Methods 2011, 54, 351-360.
-
Giansanti, P.; Tsiatsiani, L.; Low, T. Y.; Heck, A. J. R. “Six alternative proteases for mass spectrometry-based proteomics beyond trypsin.” Nat. Protoc. 2016, 11, 993-1006.
Guidelines, protocols, and examples are given in the following references:
- “Overview of Affinity Purification.” (Pierce/Thermo)
- Dunham, W. H.; Mullin, M.; Gingras, A.-C. “Affinity purification coupled to mass spectrometry: Basic principles and strategies.” Proteomics 2012, 12, 1576-1590.
- Pflieger, D.; Bigeard, J.; Hirt, H. “Isolation and characterization of plant protein complexes by mass spectrometry.” Proteomics 2011, 11, 1824-1833.
- Paul, F. E.; Hosp, F.; Selbach, M. “Analyzing protein-protein interactions by quantitative mass spectrometry.” Methods 2011, 54, 387-395.
- Malovannaya, A.; Li, Y.; Bulynko, Y.; Jung, S. Y.; Wang, Y.; Lanz, R. B.; O’Malley, B. W.; Qin, J. “Streamlined analysis schema for high-throughput identification of endogenous protein complexes.” Proc. Natl. Acad. Sci. USA 2010, 107, 2431-2436.
For a detailed oligonucleotide sample preparation protocol, please see Shah, S.; Friedman, S. H. “An ESI-MS method for characterization of native and modified oligonucleotides used for RNA interference and other biological applications.” Nature Protocols 2008, 3, 351-356.
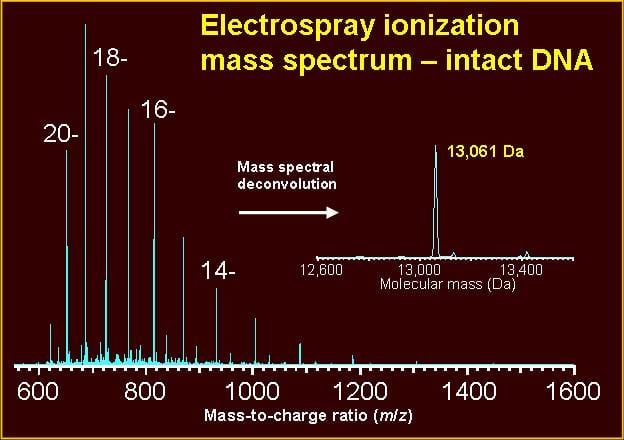
Guidelines, protocols, and examples are given in the following references:
- Lorenzen, K.; Duijn, E. “Native mass spectrometry as a tool in structural biology.” Current Protocols in Protein Science 2010, 62, 17.12:17.12.1-17.12.17.
- Heck, A. J. R. “Native mass spectrometry: A bridge between interactomics and structural biology.” Nature Methods 2008, 5, 927-933.
- Mehmood, S.; Allison, T. M.; Robinson, C. V. “Mass spectrometry of protein complexes: From origins to applications.” Annu. Rev. Phys. Chem. 2015, 66, 453-474.
- Liu, T. Y.; Iavarone, A. T.; Doudna, J. A. “RNA and DNA targeting by a reconstituted Thermus thermophilus Type III-A CRISPR-Cas system.” PLoS ONE 2017, 12, e0170552.
Examples are given in the following references:
- Offenbacher, A. R.; Hu, S.; Poss, E. M.; Carr, C. A. M.; Scouras, A. D.; Prigozhin, D.; Iavarone, A. T.; Palla, A.; Alber, T.; Fraser, J. S.; Klinman, J. P. “Hydrogen-deuterium exchange of lipoxygenase uncovers a relationship between distal, solvent exposed protein motions and the thermal activation barrier for catalytic proton-coupled electron tunneling.” ACS Cent. Sci. 2017, 3, 570-579.
- Underbakke, E. S.; Iavarone, A. T.; Marletta, M. A. “Higher-order interactions bridge the nitric oxide receptor and catalytic domains of soluble guanylate cyclase.” Proc. Natl. Acad. Sci. USA 2013, 110, 6777-6782.
- Tsutsui, Y.; Wintrode, P. L. “Hydrogen/deuterium exchange-mass spectrometry: A powerful tool for probing protein structure, dynamics and interactions.” Current Medicinal Chemistry 2007, 14, 2344-2358.